Looking Out for the Little Guys

A new study, undertaken by a team of scientists from BIOS, the Bermuda Government Department of Environment and Natural Resources, and the University of Rhode Island, used DNA markers to characterize the genetic diversity of Bermuda’s baitfish populations. The island’s baitfish, as seen here in a large assemblage referred to as a “bait ball,” were shown to exist in highly mixed populations of up to six different species. Photo credit: Joanna Pitt.
Small fishes play an important role in the marine food chain, providing food for larger fishes and water birds, but they are also caught for use as bait in both commercial and recreational fisheries. Over the past thirty years, a decline has been noted in some species of baitfish, leading scientists and resource managers to look more closely at the population dynamics of these important fish. However, baitfish tend to congregate in large schools containing multiple species, making it difficult to study individual populations.
In 2017, a team of researchers, with funding from a Darwin Plus grant through the United Kingdom’s Department of Environment, Food and Rural Affairs (DEFRA), set out to characterize Bermuda’s baitfish populations, with the goal of providing data to improve the management of these species. Currently, regulations are in place that restrict the size of nets that may be used to catch baitfish, while net fishing is prohibited altogether in four inshore bays around the island: Coot Pond, Whalebone Bay, and Shelly Bay in the east end, and Somerset Long Bay in the west end. However, one important question remaining for the research team was whether similar protected bays need to be more evenly distributed around the island.
Gretchen Goodbody-Gringley, a coral reef ecologist at BIOS, and Joanna Pitt, marine resources officer for the Bermuda Government Department of Environment and Natural Resources, led the project, which also included Emma Strand, a former BIOS Research Experiences for Undergraduates (REU) student and a current doctoral student at the University of Rhode Island. Their findings were recently published as a paper in the peer-reviewed scientific journal PeerJ.
“Baitfish depend on shallow inshore areas that are often modified and threatened by a variety of human activities, such as overfishing, pollution, and coastal development,” Strand said. “It’s important to understand the composition of baitfish assemblages and the distribution of the various species around the island.”
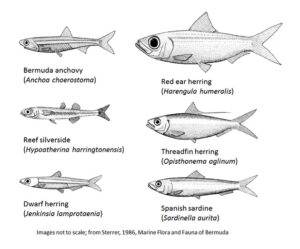
The six most common baitfish species in Bermuda, shown here as line drawings, were found to have a high degree of genetic diversity both within and between assemblages, indicating that individual fish from multiple locations around the island contribute to a single gene pool. The study’s results also indicate that Bermuda’s baitfish populations are largely locally maintained and not likely to be replenished from populations in the Caribbean or other areas of the Tropical West Atlantic. Illustration credit Smith-Vaniz et al. (1999)
During June and July of 2017, the team collected baitfish from ten locations around Bermuda, including both the east and west ends and north and south shores. Using physical characteristics, they identified five of the most common baitfish species in Bermuda: the reef silverside (Hypoatherina harringtonensis), the Bermuda anchovy (Anchoa choerostoma), the dwarf herring (Jenkinsia lamprotaenia), the red ear herring (Harengula humeralis), and the round sardinella or Spanish sardine (Sardinella aurita).
DNA samples were extracted from the muscle of the collected fish and then sequenced, allowing the researchers to look at the genetic makeup of the baitfish populations. The team looked at specific DNA markers found in mitochondria, which are the parts of a cell responsible for converting food into energy. Analysis of mitochondrial DNA helps shed light on how genetically diverse each fish species is at the local level, as well as how genetically connected (or similar) Bermuda’s local populations are to populations elsewhere. It also serves as a method for confirming the initial identification of each species, which increases the accuracy of the results.
Based on these genetic analyses, the team realized that the samples included a sixth species, the Atlantic thread, or threadfin, herring (Opisthonema oglinum), highlighting the importance of conducting DNA analyses when characterizing multispecies baitfish assemblages.
“For all six of the species we examined, our results showed a high degree of genetic diversity both within and between baitfish assemblages from different locations around Bermuda,” Goodbody-Gringley said. “This tells us that, in terms of management, we should consider Bermuda’s baitfish species as highly mixed populations in which individuals from all around the island contribute to a single gene pool.”
However, the results also indicated that gene flow, and therefore exchange of individuals, between populations of these species in Bermuda and those in other regions of the Caribbean and Western Atlantic is limited, suggesting that the Bermuda populations are largely locally maintained. From a management perspective, this means that local baitfish populations are not likely to be replenished from populations in the Caribbean or other areas of the Tropical West Atlantic.
“As a resource manager, it’s important to be able to balance the ecological sustainability of baitfish populations and their role in commercial and recreational fishing,” Pitt said. “In practical terms, these two pieces of information mean that we need to work to conserve our baitfish stocks, but that we don’t need to worry about the exact placement of protected areas.”
She added that studies such as this demonstrate the value of population genetics for informing and adapting fishery management measures.
